Chemotaxis, introduced in my post on January 26, is the process by which cells migrate along a gradient of growth factor. Researchers study chemotaxis for many different reasons. First, there’s the biological study of how these cells move. Then, there’s the study of why these cells move. In my post from September 28, I describe how random movement of cells in response to an even distribution of growth factor can be measured. However, the next step is to measure whether this response can be seen in a directional manner in response to a gradient.
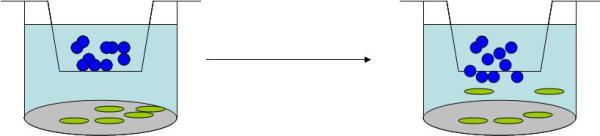
The easiest method, perhaps, for studying chemotaxis is the Boyden Chamber assay. The concept of the Boyden Chamber assay is straightforward. Known as a transwell system, a smaller well is suspended over the main well of a tissue culture plate. However, the bottom of this insert is not solid plastic, put a permeable membrane. Cells are added to the top of this membrane, in solution, and their migration across the membrane to the other side in response to stimuli can be measured simply by counting the cells that have passed through. A great product for this are the FluoroblokTM inserts from BD Biosciences. Cells can be fluorescently labeled, and light from cells on the top side of the membrane won’t shine through to the bottom. The layout of a basic Boyden Chamber Assay is shown (blue dots are cells, green dots are stimuli):
The Boyden Chamber Assay may be simple, but requires a high level of finesse. The first main concern is the use of proper controls. Not only are negative and positive controls important, but also we need to measure the directionality of response to control for random movement. I.e., although, for example, growth factor might be added to the bottom well, and cells to the top, we need to show that the a gradient is actually maintained and that it is not simply a case of growth factor being evenly distributed on either side of the membrane (equilibrium) and cell movement increased in general.
Also, there are several steps in which human inaccuracy can lead to deviation between wells, ultimately resulting in lack of statistical significance because of too much variability. The assay is indeed incredibly sensitive to both the number of cells and the amount of stimulus, both of which tend to be very low, meaning that everything must be incredibly accurate. The first year of my PhD I conducted a gigantic Boyden Chamber assay, but got no significant results, just trends, because the error bars were too large.
The third part of the assay which requires delicacy is the counting of the cells. As I mentioned, the cells are fluorescently stained. However, in order to calculate fluorescence, because the desired cells are on the bottom of the membrane, in order to use spectrophotometry, you’d have to have a bottom plate reader. Our spectrophotometer could not be reprogrammed to do this (although in theory it had the ability). This meant that the membrane had to be cut off and mounted upright before being imaged. The cutting of the membrane is tricky, because it becomes easy to crease, which can lead to problems when trying to mount it flat (it’s very difficult to image a crinkled sample due to focus issues).
The protocol for Boyden Chamber Assays:
- Trypsin cells, centrifuge, resuspend in desired medium at desired density
- Make sure this cell suspension is evenly mixed
- Place cells in the top of the transwell
- Place desired medium in the bottom chamber
- Make sure the volume of the bottom corresponds to the top volume so no fluid pressure is exerted across the membrane (manufacturer’s recommendations)
- Incubate for the desired amount of time (good idea to perform a time-course with controls as a preliminary study)
- Gently aspirate medium on top and bottom, wash 3x with PBS
- Incubate 15 minutes in 4% PFA in PBS in order to fix the cells
- Wash 3x with PBS
- Incubate with 1:10000 DAPI in TBS with 0.1% Tween for 5 minutes at 37°C.
- Wash 3x with PBS
- Carefully cut out membranes and place, bottom-up on slides
- Mount with fluorescence-stabilizing hard-set mounting medium
- Pressing down to eliminate any air bubbles and make sure membrane is flat
- Image at 4x magnification
- Use ImageJ to increase brightness/contrast to max, and convert to binary for quantification